Shimadzu has produced an application note describing phospholipid analysis for four types of mouse tissues performed by precursor ion scan (PIS) or neutral loss scan (NLS) with a triple quadrupole mass spectrometer.
 Introduction
Introduction
Phospholipids are a class of biologically important molecules that share the same basic structure – a head group linked to fatty acid tail group – with high diversity caused by various combinations of head and tail. They play a major role in the formation of lipid bilayer as a major component of the cell membrane together with glycolipids and cholesterol in vivo. In addition to constituting cell membranes, phospholipids also function as a source of fatty acids involved in energy production by lipid metabolism, lipid transport and signal transduction. Thus, although a phospholipid works for various functions in vivo, its number is enormous because of the combination of a characteristic head group and fatty acid constituting its structure. Therefore, in the field of lipidomics, qualitative analysis utilizing an enormous database is often performed based on information such as m/z of phospholipids, the product ion derived from the characteristic head group and from each fatty acid.
 Method
Method
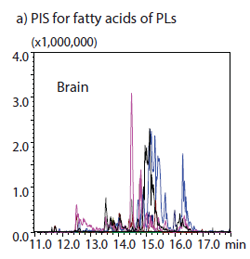
In this application note analysis was performed by precursor ion scan (PIS) or neutral loss scan (NLS) with a triple quadrupole mass spectrometer to compare the phospholipid profiles expressed by tissues of mice brain, spleen, lung and liver. Peaks detected by PIS and NLS were assigned to candidate phospholipids by the database search using SimLipid software from PREMIER Biosoft, USA (www.premierbiosoft.com).
Results
Phospholipids that are known to be characteristic of each tissue were correctly detected from four different tissue extracts, confirming the effectiveness of the database search by SimLipid software.




